National Grants
DIAMARK
Project code: PN-III-P4-PCE-2021-1344
Contract number: PCE 114/2022
Project title:
Molecular markers for prediction of evolution and prognosis in diabetic foot management
Markeri moleculari ca predictori ai evolutiei si prognosticului in managenentul piciorului diabetic (DFS)
Acronym: DIAMARK
Project duration: 02/06/2022 - 31/12/2024
Project director: Felicia Antohe, PhD, CS I
Project team:
- Ivan Luminita, PhD, CSII
- Uyy Elena, PhD, CS III
- Suica Viorel Iulian, PhD, CSIII
- Boteanu Raluca Maria, PhD, CSIII
- Chiriac Ovidiu, MD
- Georgescu Dragos Eugen, MD
- Bernea Georgiana Elena, ex-member, PhD Student
- Uta Valentina Diana, PhD Student
Abstract: Chronic hyperglycaemia has a life-threatening part in the pathogenesis of peripheral artery disease (PAD), leading to vascular complications by means of metabolic and structural abnormalities. The proposal aims to reveal correlations in subjects with diabetic foot syndrome (DFS) between immuno-inflammatory damage associated molecular patterns (DAMPs) and clinical and laboratory parameters, as a confirmation of the crucial role of DAMPs in PAD. Proteomics evaluation of potential markers in DFS could offer significant insights in the pathogenesis of ulcers and their micro and macro-vascular background, offering a predictive role of foot complications and stratification. A DAMP panel identified and quantified in the plasma of DFS patients will improve the unsolved yet clinical resolution of inflammatory process associated to the early events in the diabetic foot ulceration. Preclinical manipulation of the innate immune response, by targeting the DAMP-receptor interaction and tissue recovery in experimental animals by controlled neutrophil extracellular traps (NETosis) cellular death, represents a potential novel, original therapeutic opportunity to reduce the hyperinflammation and tissue damage associated with the chronic wounds of DFS, and to restart the normal wound healing process. Consequently, we can foresee that certain DAMPs can be envisioned as biomarkers for the evolution and prognosis of DFS but also as molecular targets in the treatment of diabetic foot ulcerations.
Rezumat: Hiperglicemia cronică are efecte devastatoare în patologia bolilor arteriale periferice, caracterizate de complicații vasculare metabolice și structurale. Scopul proiectului este de a stabili corelații între modificările proteomice ale expresiei moleculelor de stres (DAMPs) și parametrii clinici prezenți în inflamația piciorului diabetic (DFS). Analiza proteomică bazata pe spectrometrie de masa a markerilor moleculari exprimați în inflamația piciorului diabetic vor conduce la înțelegerea patologiei micro- și macro-vasculare și vor putea fi folosiți în predicția și stratificarea pacienților cu boli arteriale periferice. Selecția și cuantificarea unui panel de biomarkeri plasmatici ar reprezenta o rezolvare clinica mult așteptată pentru preventia, diagnosticarea timpurie si strategia adecvata de tratament a inflamației piciorului diabetic. Manipularea preclinică a răspunsului imun înnăscut, prin țintirea interacțiunii receptori-proteine de stres si reducerea morții celulare controlate prin NEToza, în animale mici de laborator, reprezintă o potențială oportunitate terapeutică nouă, originală cu scopul de a reduce hiperinflamația și leziunile tisulare asociate cu rănile cronice ale DFS și pentru a relua procesul normal de vindecare a rănilor. În consecință, putem prevedea că anumite DAMPs pot fi biomarkeri valoroși pentru evoluția și prognosticul DFS, dar și ținte moleculare în tratamentul ulcerațiilor piciorului diabetic.
Objectives:
O1: Identification of DAMPs from the peripheral arteries of DFS patients.
O2: Quantitative correlation between DAMPs identified in the DFS peripheral arteries and plasma of patients with different stages of diabetic foot evolution.
O3: Evaluation of a therapeutic strategy on a laboratory animal model of diabetic foot using a targeted approach against DAMP-generated inflammatory response and NETosis.
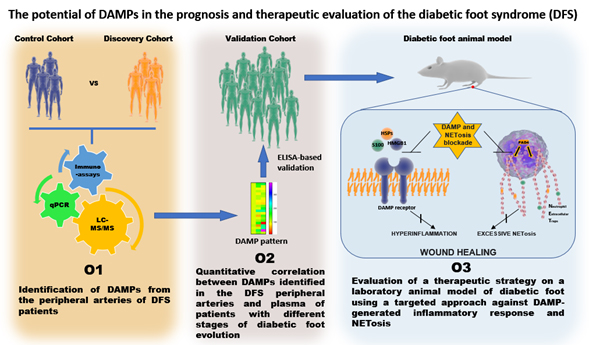
Figure 1. Graphical abstract. The proposed workflow of the project
Report: 2022
O1: Identification of DAMPs from the peripheral arteries of DFS patients.
Activity 1.1: Professional patients’ enrolment. Clinical and paraclinical profiling. Plasma sample collection. Patient follow-up.
Activity A1.2: Texas wound classification and PAD status evaluation. Peripheral small artery isolation. Texas wound classification was used to quantify severity of the lesions.
Activity A1.3: Histological assessment of discovery group (DG) peripheral small arteries.
Activity A1.4: Quantitative profiling of DAMPs from the peripheral arteries of DG vs CG samples.
LC-MS/MS experiments were performed using the Easy Nano-LC II system (Thermo Scientific, San Jose, CA) coupled with the LTQ - Velos Pro Orbitrap ETD mass spectrometer (Thermo Scientific, San Jose, CA, USA). The high-performance LTQ-Velos Orbitrap system was used to generate mass spectra using the Top 15 Data-dependent method. Protein identification was performed using Proteome Discoverer 2.4 (Thermo Scientific, San Jose, CA). Oxidation of methionine and deamidation of asparagine and glutamine were established as dynamic modifications while carbamidomethylation at cysteine was established as a fixed modification.
Chromatographic elution optimization of peptide mixtures was performed, with an acetonitrile gradient of 3-25% over 90 min demonstrating optimal separation yield of both hydrophilic and hydrophobic peptides (Figure 2).
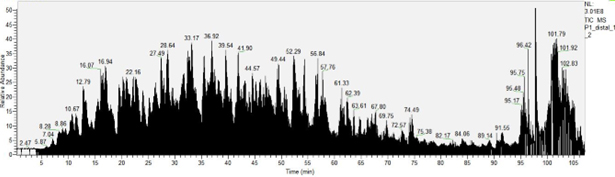
Figure 2: Chromatographic separation of the complex mixture of peptides from a representative sample. A 3-25% gradient of acetonitrile in water was applied for chromatographic separation of peptides using Pep-Map analytical column (Thermo Scientific) with the following characteristics: 15 cm length, 75 µm inner diameter, C18 chain, 3 µm sphere size, 120 Å port diameter.
The raw mass spectra data were processed using the Proteome Discoverer 2.4 software bioinformatic platform. This resulted in the identification of 1330 proteins, out of which 442 were found in all 3 samples (Figure 3). In order to increase the confidence of protein identification, 2 sets of filters were applied, at the level of Sequest score (>10), and protein false positive discovery rate (FDR<0.05).
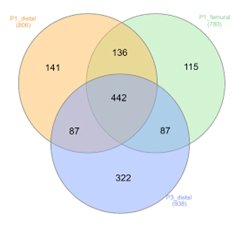
Figure 3: Venn intersection of the identified proteins.
Report: 2023
O2: Quantitative correlation between DAMPs identified in the DFS peripheral arteries and plasma of patients with different stages of diabetic foot evolution.
O3: Evaluation of a therapeutic strategy on a laboratory animal model of diabetic foot using a targeted approach against DAMP-generated inflammatory response and NETosis.
Chronic hyperglycemia has devastating effects in the pathology of peripheral arterial disease (PAD), characterized by metabolic and structural vascular complications. In this context, the identification of new inflammatory markers and the association with an increased risk of death or amputation or other events in the lower limbs in the early stage of the disease is necessary. Analysis of inflammatory biomarkers is useful to achieve risk stratification in PAD at different stages of evolution. This phase continued with the collection of tissue and blood samples and the addition of new recruits to the control (GC), validation (GV) and investigation (GI) groups. Also in this phase, the classification of the wounds was completed and an evaluation of the pathology of peripheral arterial diseases was carried out, including through immunohistochemistry analysis through other types of staining (Masson and van Kossa trichrome).
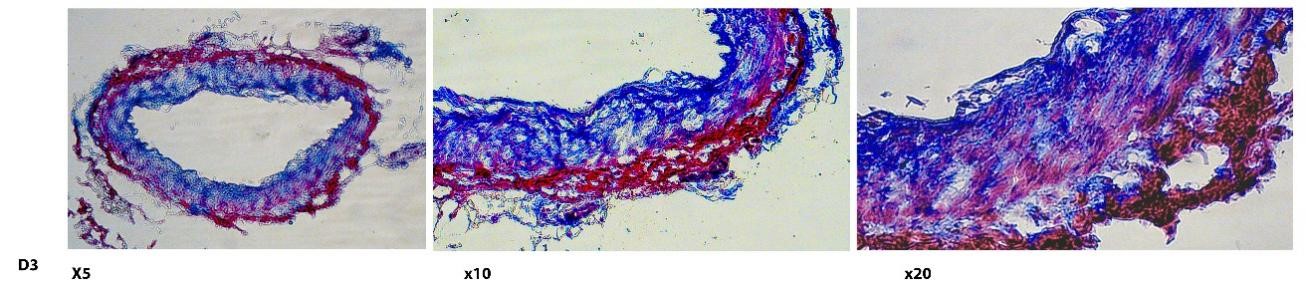
Figure 4: Fibrosis in the pedis artery. Trichrome Masson staining demonstrates total collagen content (blue).
In this activity, the acquisition and construction of the library of spectral data necessary to extract the qualitative and quantitative information for the molecules of interest (DAMP) continued. The quantitative profile of the 21 DAMPs identified in the peripheral arteries of DFS patients and 15 molecules from plasma patients were carried out. DAMPs that change significantly in the plasma of patients under the action of vascular pathology were evaluated. Statistically over-represented KEGG (Kyoto Encyclopedia of Genes and Genomes) signaling pathways were also identified. The degree of Pearson correlation was analyzed between the spectral abundances of 5 commune DAMP proteins that change their expression both in plasma and in excised vascular tissue from patients with diabetes and DFS (GI group). Experiments were also performed on a mouse diabetic model with ischemia and leg wound that closely mimic the human ppathology.
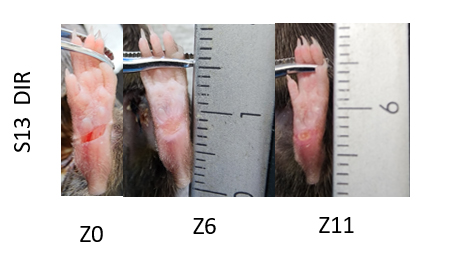
Figure 5: Evolution of the leg injury over time. Image of the paw wound in a mouse (S13) with diabetes and femoral artery ischemia on days 0 (immediately after surgery), 6 and 11 of the experiment. DIR: diabetes, ischemia and wound.
The evaluation of the animal model will be performed in the next stages of the project.
Report: 2024
Objectives:
O1: Identification of DAMPs from the peripheral arteries of DFS patients.
O2: Quantitative correlation between DAMPs identified in the DFS peripheral arteries and plasma of patients with different stages of diabetic foot evolution.
O3: Evaluation of a therapeutic strategy on a laboratory animal model of diabetic foot using a targeted approach against DAMP-generated inflammatory response and NETosis.
O1 and O2: Following mass spectrometry analysis of human plasma, 15 DAMP molecules were identified that show a regulation of at least 1.2-fold in the plasma of patients, that is induced by diabetes alone or diabetes with DFS, and 21 DAMP proteins in excised vascular tissue with altered abundance in the different stages diabetic foot ulceration. The intersection of the two protein lists led to the identification of 5 common proteins: fibrinogen beta chain (FGB), fibrinogen gamma chain (FGG), neutrophil defensin 1 (DEFA1B), serum amyloid A-1 protein (SAA1), tenascin-X (TNXB).

Figure 6: The illustrations show the intersection of DAMP proteins whose spectral abundance is altered 1.2-fold in plasma/vascular tissue collected from patients with diabetes and DFS versus controls (A) and the Pearson correlation analysis between DAMP proteins common to the two types of samples (B). The Pearson correlation coefficient is presented from 1 (red with the maximum degree of positive correlation) to -1 (blue with the maximum degree of negative correlation).
O3: Proteomic analysis identified in the plasma of the animals the proteins involved in the inflammatory response and tissue remodeling in the context of ischemia and the treatments applied.

Figure 7: Heatmap illustrating significant variations in plasma inflammatory markers, with an abundance change of at least 1.2-fold, associated with ischemia and/or the type of treatment administered. The analysis was performed on plasma samples collected from the experimental groups: 1. control with diabetes and plantar lesions (DR), 2. diabetes with ischemic and plantar lesions (DIR), 3. diabetes with ischemic and plantar lesions treated with TAK 242 (DIRT1), 4. diabetes with ischemic and plantar lesions treated with the NETosis inhibitor (DIRT2), and 5. diabetes with ischemic and plantar lesions treated concomitantly with both therapies, T1 and T2 (DIRT1T2). Markers with changes in abundance were considered significant at an adjusted p-value < 0.05.
The proteins identified in the plasma of animals were integrated in the KEGG databases, and these proteins were associated with 11 over-represented signaling pathways, such as Lysosome (FDR =4.10e-03), Phagosome (FDR=4.10e-03), Apoptosis (FDR =4.10e-03), Malaria (FDR =4.10e-03), Focal adhesion (FDR =4.20e-03), Salivary secretion (FDR =5.20e-03), ECM-receptor interaction (FDR=5.90e-03), Autophagy – animal (FDR =1.92e-02), PI3K-Akt signaling pathway (FDR=2.120e-02), Ferroptosis (FDR =3.21e-02), Mineral absorption (FDR=4.64e-02), highlighting the influence of treatments on key biological processes in the ischemic diabetic foot.

Figure 8: Statistical over-representation analysis of KEGG signaling pathways involving DAMPs identified by mass spectrometry that show at least a 1.2-fold change in their abundance induced by ischemia and/or the type of treatment administered. Pathways are ordered according to statistical significance FDR: False discovery rate.
Performance indicators 2022 – 2024:
- 4 ISI papers with a cumulative impact factor of 13.835 + 11.4 (in press) = 25.235;
- 4 international ISI-indexed databases:
1. PRIDE PXD033683 - Short-term blockade of pro-inflammatory alarmin S100A9 favorably modulates left ventricle proteome and related signaling pathways involved in post-myocardial infarction recovery
Public release: 2022-05-19
Author: Felicia Antohe
Repository: PRIDE (Proteomics Identification Database) 2022
2. PXD035692 - Coronary alarmins as residual risk markers of atherosclerosis under hypolipidemic therapy
Public release: 2022-10-03
Author: Felicia Antohe
Repository: PRIDE (Proteomics Identification Database) 2022
3. PXD035673 - Exosome proteomics reveals the deregulation of coagulation, complement, and lipid metabolism proteins in gestational diabetes mellitus
Public release: 2022-09-22
Author: Felicia Antohe
Repository: PRIDE (Proteomics Identification Database) 2022
4. PXD026379 - Alteration of DAMPs and regulated cell death under hyperlipidemic stress
Public release: 2022-02-22
Author: Felicia Antohe
Repository: PRIDE (Proteomics Identification Database) 2022
- 2 oral presentations at international conferences;
- 14 posters presented at international scientific events:
- 5 posters (2022), 3 posters (2023), and 6 posters (2024).
- 2 poster awards (awarded to PhD student Diana Uta and Dr. Viorel Suica);
- 1 article awarded by UEFISCDI:
PN-IV-P2-2.3-PRECISI2023-71403 Șuică Viorel et al. - Cardiac Alarmins as Residual Risk Markers of Atherosclerosis under Hypolipidemic Therapy - 2022

